pacman::p_load(igraph,tidygraph, ggraph, visNetwork,lubridate,clock,tidyverse,graphlayouts)In class exercise network
Load packages:
Importing network data from files
GAStech_nodes <- read_csv("data/GAStech_email_node.csv")
GAStech_edges <- read_csv("data/GAStech_email_edge-v2.csv")create 2 more columns
GAStech_edges <- GAStech_edges %>%
mutate(SendDate = dmy(SentDate)) %>%
mutate(Weekday = wday(SentDate,
label = TRUE,
abbr = FALSE))Wrangling attributes
A close examination of GAStech_edges data.frame reveals that it consists of individual e-mail flow records. This is not very useful for visualisation.
In view of this, we will aggregate the individual by date, senders, receivers, main subject and day of the week.
GAStech_edges_aggregated <- GAStech_edges %>%
filter(MainSubject == "Work related") %>%
group_by(source, target, Weekday) %>%
summarise(Weight = n()) %>%
filter(source!=target) %>%
filter(Weight > 1) %>%
ungroup()Using tbl_graph() to build tidygraph data model.
In this section, you will use tbl_graph() of tinygraph package to build an tidygraph’s network graph data.frame.
Before typing the codes, you are recommended to review to reference guide of tbl_graph()
GAStech_graph <- tbl_graph(nodes = GAStech_nodes, edges = GAStech_edges_aggregated, directed = TRUE)GAStech_graph %>%
activate(edges) %>%
arrange(desc(Weight))# A tbl_graph: 54 nodes and 1372 edges
#
# A directed multigraph with 1 component
#
# Edge Data: 1,372 × 4 (active)
from to Weekday Weight
<int> <int> <ord> <int>
1 40 41 Saturday 13
2 41 43 Monday 11
3 35 31 Tuesday 10
4 40 41 Monday 10
5 40 43 Monday 10
6 36 32 Sunday 9
7 40 43 Saturday 9
8 41 40 Monday 9
9 19 15 Wednesday 8
10 35 38 Tuesday 8
# ℹ 1,362 more rows
#
# Node Data: 54 × 4
id label Department Title
<dbl> <chr> <chr> <chr>
1 1 Mat.Bramar Administration Assistant to CEO
2 2 Anda.Ribera Administration Assistant to CFO
3 3 Rachel.Pantanal Administration Assistant to CIO
# ℹ 51 more rowsPlotting a basic network graph
The code chunk below uses ggraph(), geom-edge_link() and geom_node_point() to plot a network graph by using GAStech_graph. Before your get started, it is advisable to read their respective reference guide at least once.
ggraph(GAStech_graph) +
geom_edge_link() +
geom_node_point()
Fruchterman and Reingold layout
The code chunks below will be used to plot the network graph using Fruchterman and Reingold layout.
g <- ggraph(GAStech_graph,
layout = "fr") +
geom_edge_link(aes()) +
geom_node_point(aes())
g + theme_graph()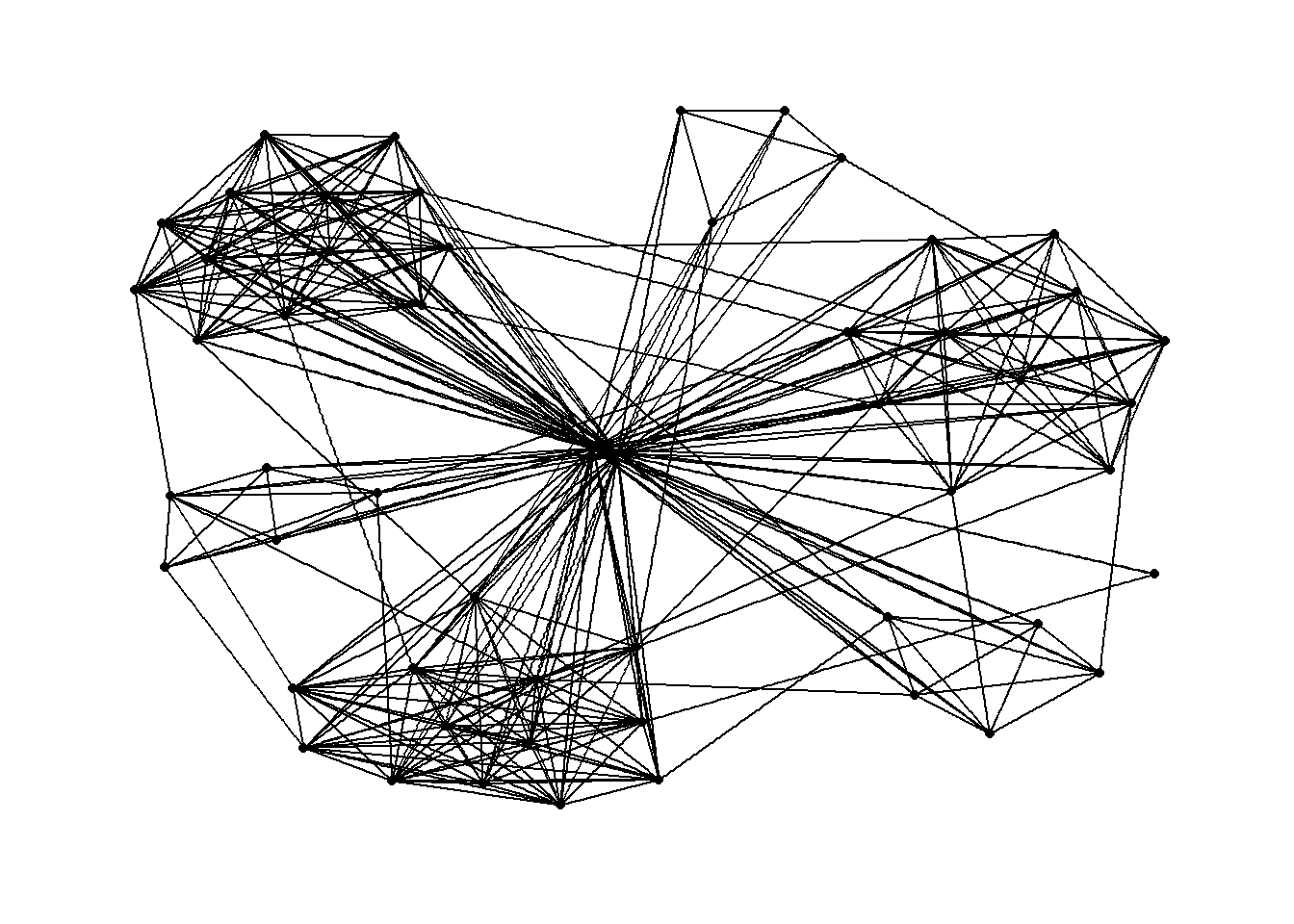
Modifying network nodes
In this section, you will colour each node by referring to their respective departments.
g <- ggraph(GAStech_graph,
layout = "nicely") +
geom_edge_link(aes()) +
geom_node_point(aes(colour = Department,
size = 3))
g + theme_graph()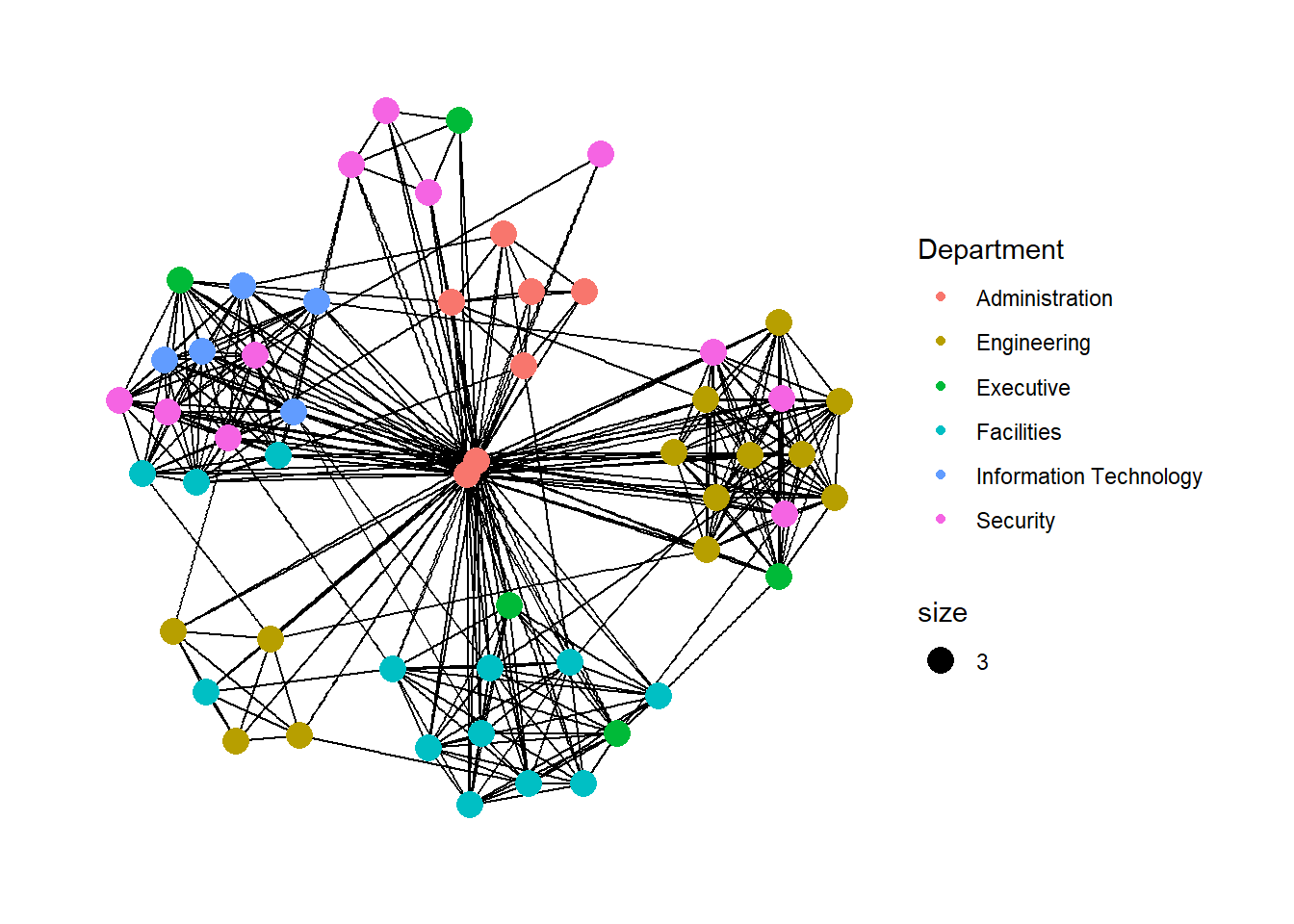
Modifying edges
In the code chunk below, the thickness of the edges will be mapped with the Weight variable.
g <- ggraph(GAStech_graph,
layout = "nicely") +
geom_edge_link(aes(width=Weight),
alpha=0.2) +
scale_edge_width(range = c(0.1, 5)) +
geom_node_point(aes(colour = Department),
size = 3)
g + theme_graph()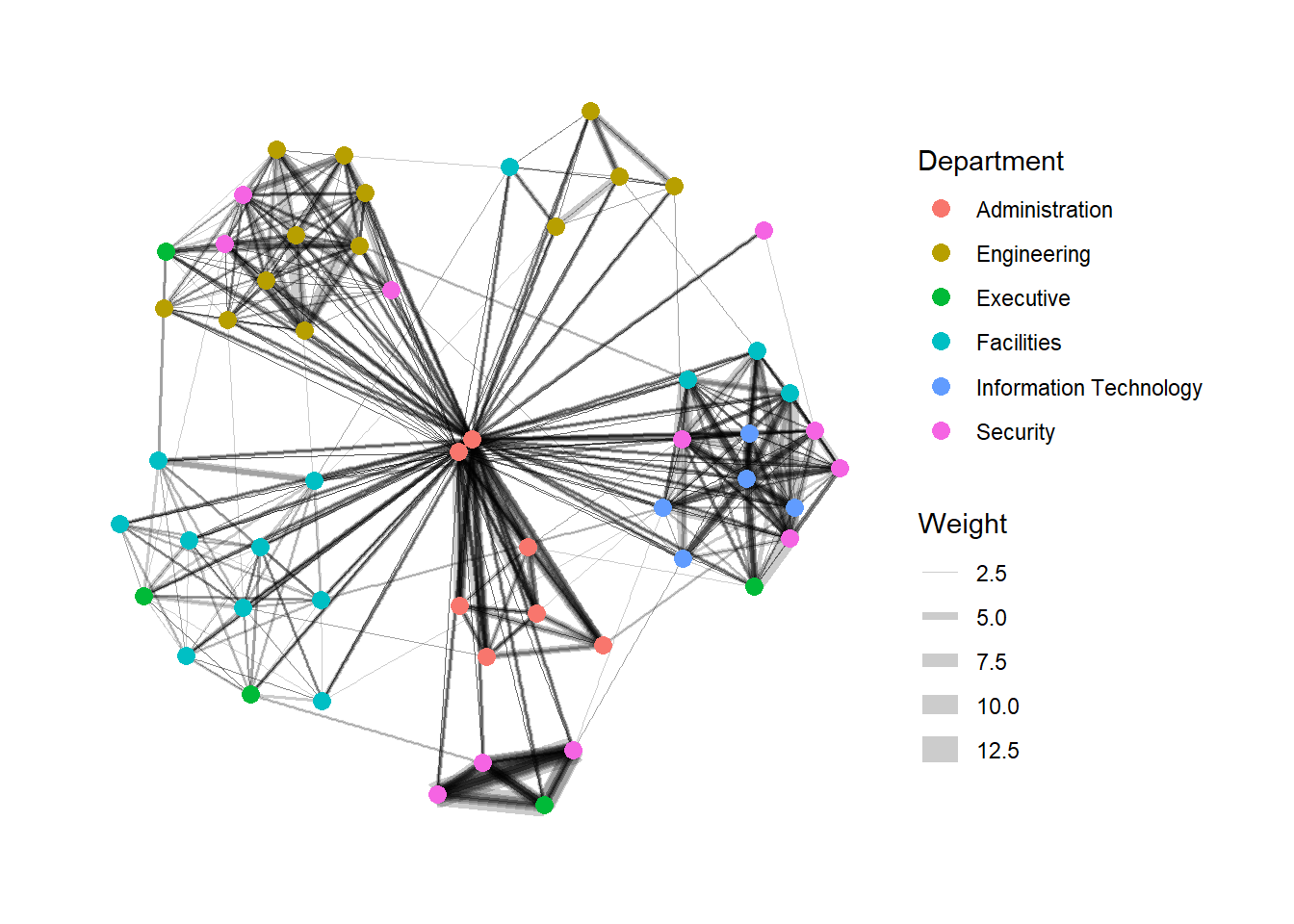
Working with facet_edges()
In the code chunk below, facet_edges() is used. Before getting started, it is advisable for you to read it’s reference guide at least once.
set_graph_style()
g <- ggraph(GAStech_graph,
layout = "nicely") +
geom_edge_link(aes(width=Weight),
alpha=0.2) +
scale_edge_width(range = c(0.1, 5)) +
geom_node_point(aes(colour = Department),
size = 2)
g + facet_edges(~Weekday)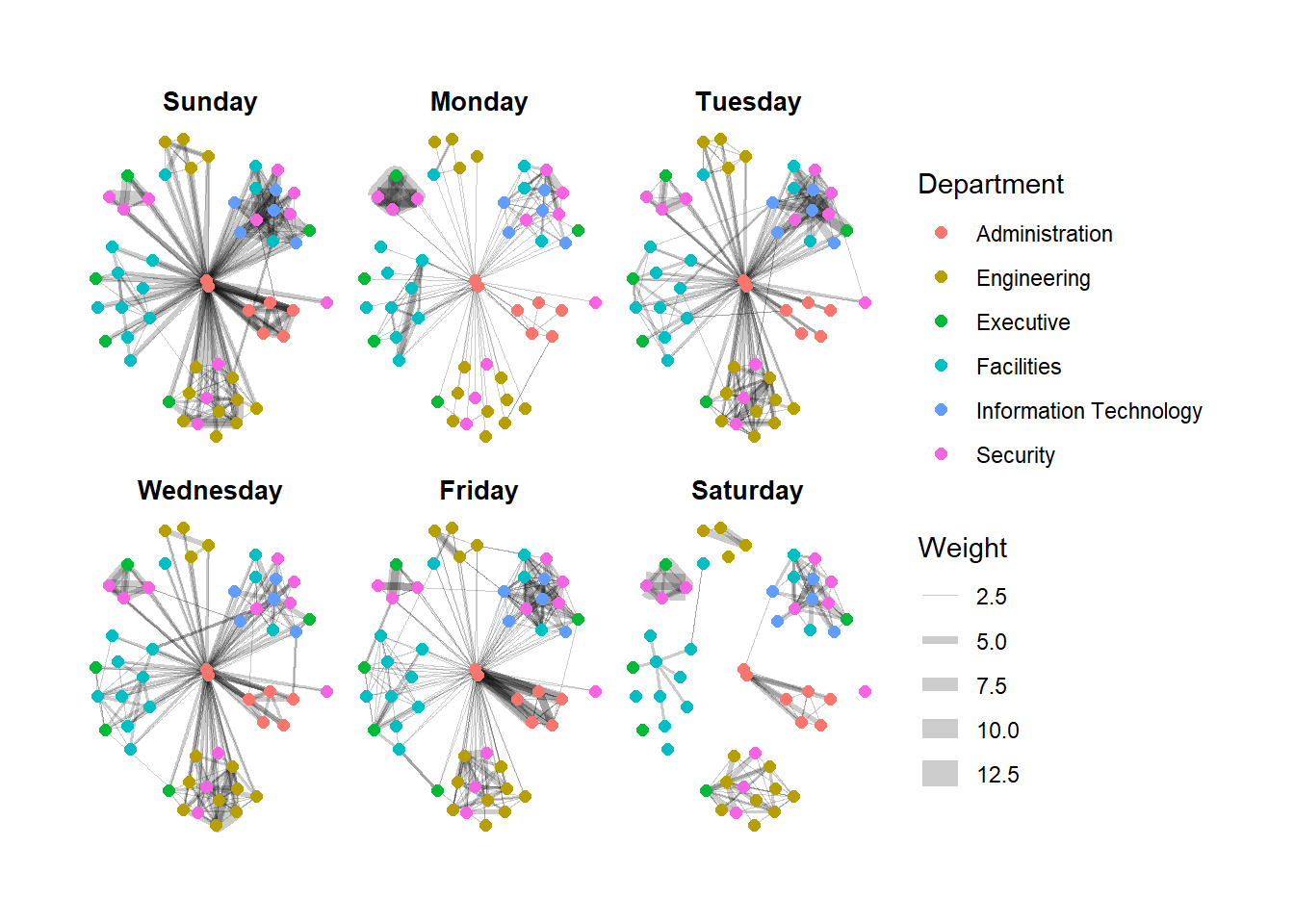
A framed facet graph
The code chunk below adds frame to each graph.
set_graph_style()
g <- ggraph(GAStech_graph,
layout = "nicely") +
geom_edge_link(aes(width=Weight),
alpha=0.2) +
scale_edge_width(range = c(0.1, 5)) +
geom_node_point(aes(colour = Department),
size = 2)
g + facet_edges(~Weekday) +
th_foreground(foreground = "grey80",
border = TRUE) +
theme(legend.position = 'bottom')